Simulations Done Right
How to do Simulations
Top Tips
🎯 Thoughtful Goals = Direction!
- practical utility
- key insight
- audience
📊 Communication = Focus!
- structure + organize
- minimize cognitive load
⚙️ Clean, Fast Sims = Productivity!
- simple DGPs
- minimize compute
- modular code
🎯 Setting Goals
Setting Goals
- paper = claims 🎯 + evidence
- evidence = simulations 📊, applications 🗂️, theory ♾️, etc.
- A good claim likely…
- has practical utility
- builds insight
- is honest and well-scoped
- Many claims are possible. Less is more! Prioritize:
- relevance to audience
- potential to change practice or thinking
- sims often support claims like: “[method] works [well] when [condition] and not when [other condition]”
Exercise: TMLE
You wake up. It’s 2006. You look in the mirror and you see you are Mark van der Laan. You’ve just invented TMLE.
- What are the claims you should set out for the TMLE paper? Which of them can be supported by theory and which by simulations?
Some Claims
- (theory) TMLE is a general method for constructing efficient estimators of pathwise differentiable estimands
- Estimating equations already does this, so while this needs to be shown, it’s not enough to prove TMLE is useful!
- TMLE is in practice “better” than…
- Plug-in estimators? AIPW is already better
- AIPW?
❓ What is meant by “better”? When should it be better and when worse?
- It works in the real world? Computationally tractable, doesn’t require specific data types, etc. (usually addressable w/ theory and a single real data application)
Simulation Goals (Initial)
- Across the board, AIPW and TMLE behave similarly in large samples when they have access to the same learners
- TMLE is better than plugins, allowing for good coverage of Wald intervals
- There are cases where TMLE is better than AIPW in small samples
Simulation Goals (Some Questions)
- Across the board, AIPW and TMLE behave similarly in large samples when they have access to the same learners
- What learners?
- By what metric(s)?
- Like AIPW, TMLE is better than plugins, allowing for good coverage of Wald intervals
- What learners?
- What should \(n\) be here?
- Presumably using IF-based CI
- Metric?
- But is there a way to get CIs for the plugins as a naive baseline?
- There are cases where TMLE is better than AIPW in small samples
- Can we back these up with some kind of theory?
- In what DGPs?
Simulation Goals (v2)
- Across the board, AIPW and TMLE behave similarly in terms of point estimate and SE estimate in large samples when they have access to the same learners
- Like AIPW, TMLE improves the MSE of plugins in moderate sample sizes and allows for good coverage of Wald intervals based on IF SEs, even compared to the same SE put around the plugin
- TMLE is better than AIPW in small samples when the outcome is bounded (?)
📊 Presentation
Mockup Evidence for Goal 1
Across the board, AIPW and TMLE behave similarly in terms of point estimate and SE estimate in large samples when they have access to the same learners
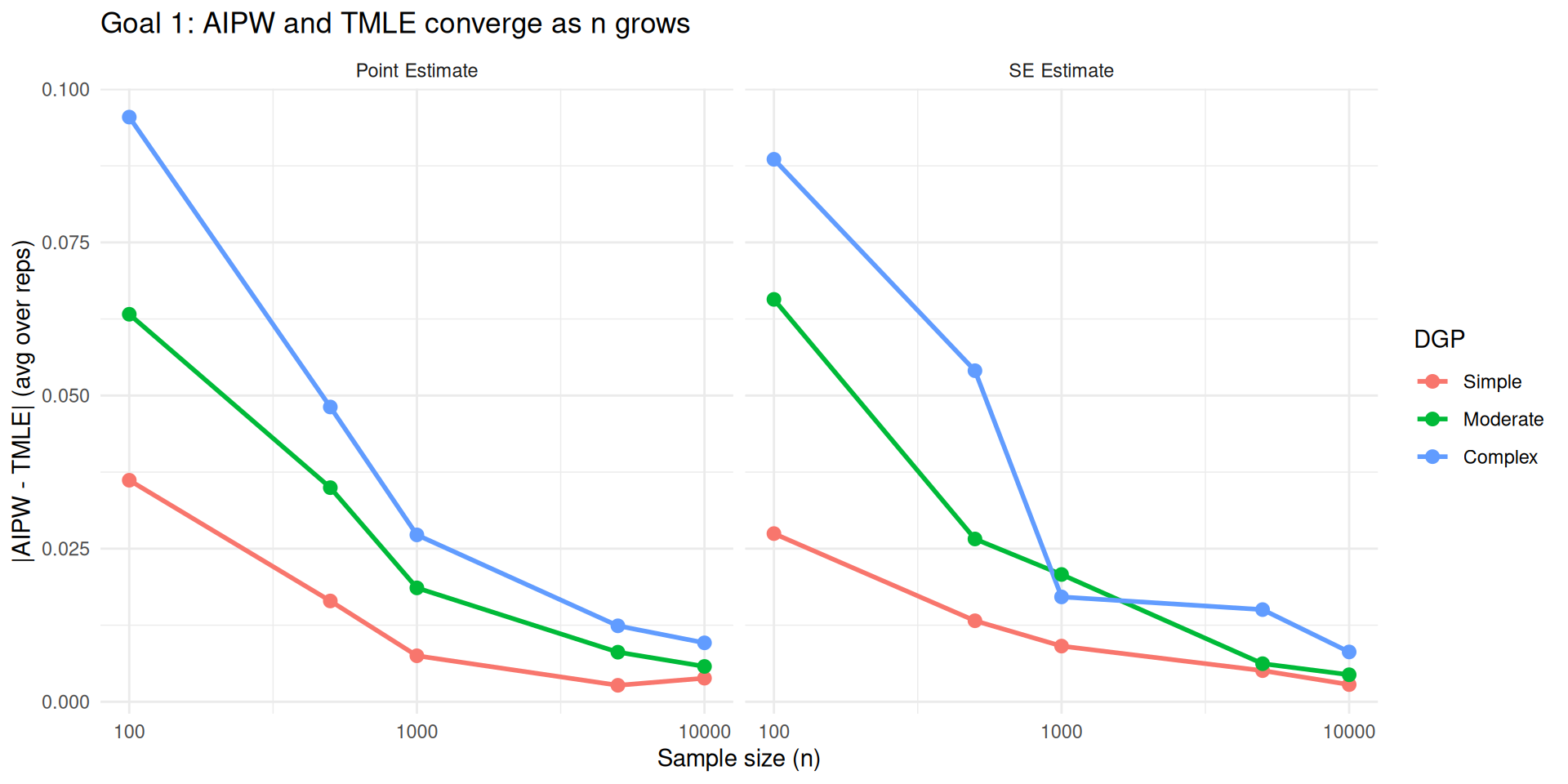
Mockup Evidence for Goal 2
Like AIPW, TMLE improves the MSE of plugins in moderate sample sizes and allows for good coverage of Wald intervals based on IF SEs.
Fix \(n = 300\), do \(B\) reps from the same 3 DGPs, tables like:
| plugin | AIPW | TMLE | |
|---|---|---|---|
| MSE | |||
| bias | |||
| variance | |||
| coverage |
One per each DGP (or 3 sub-tables). Probably just ATE, send the other estimand to the appendix.
- Learners? Some fast nonparametric thing, not too relevant to goal what it is
Mockup Evidence for Goal 3
TMLE is better than AIPW in small samples when the outcome is bounded (?)
The evidence for this goal might be a subset of evidence for the above. So just suggests that one of our DGPs should have a bounded outcome, and then we can point to the relevant rows in the above table to support this claim.
Telling the Story
For each bit of evidence you present, you should tell the reader why it is there. Tell them your goal! Then give them the evidence. Then tell them how the evidence supports the goal. Be prosaic and direct.
“In the first part I tell ’em what I am going to tell ’em; in the second part — well, I tell ’em; in the third part I tell ’em what I’ve told ’em.”
Example
The point of this application is to show how fast TMLE is on a real-world dataset of size blah blah and to prove that it works with mixed datatypes, demonstrate how it can be prespecified, and produces plausible results.
ADEMP Framework
From Morris et al. (2019)
“Using simulation studies to evaluate statistical methods”
Statistics in Medicine, 38(11), 2074–2102.
Before presenting results, clearly describe:
- Aims: what are the claims you are trying to support?
- Data-generating mechanisms: how did you simulate the data? →
DGPs - Estimands: what are the true values? → e.g.
ATE()helper - Methods: what methods are you using? →
ESTIMATORS - Performance measures: what metrics? → the
summarizestep and your tables/figures
- Aims is most important, others are in service of that
- This is part of the “tell them what you’re going to tell them.”
- Can do separately for each claim or once
Example from a Recent Paper
⚙️ Implementation
Advice for Machine Learning
- Avoid super-learner and cross-validation
- use single-split, 1-2 good learners
- Understand learners and hyperparams
ML Method Recommendations
| Method | Speed | Notes |
|---|---|---|
MARS (earth) |
⚡ Super fast | Works well with a single set of hyperparams. Ensure depth is high enough, enable pruning, allow interaction. |
| Random forests | 🏃 Fast | Works well with default hyperparams. Worse for smooth functions. |
| GBT (lightgbm/xgboost) | 🏃 Fast | Beats almost everything. Requires tuning over # trees, depth, learning rate. Use early stopping. |
| Elastic net | 🏃 Fast (ridge) | Loses to others with moderate nonlinearity. Good for baselines. Requires tuning regularization. |
| KRR (kernel ridge) | 🐌 Slow | Good for smooth functions, easy to analyze theoretically. Use only for \(n < 1000\). |
ML Method Recommendations
- Deep learning?
- Avoid unless you specifically need a differentiable model, custom loss function, fine-tuning, etc.
- Hard to tune
- xgboost wins for tabular data.
- HAL? lol. lmao even
Advice for DGPs
Being able to reason about your simulations and iterate on them is much more important than making them realistic. For example:
- \(X \sim \mathsf{Unif}([-1,1]^d)\) unless otherwise required. Bounded domain helps reason about function ranges
- \(A \sim \mathsf{Bern}(\text{expit}(\rho(X)))\). Keep \(\rho\) simple + sparse and, unless empirical positivity violations are your goal, shoot for \(\rho \in [-3,3]\)
- \(Y \sim \mu(A, X) + \mathsf{N}(0,\sigma)\). Keep \(\mu\) simple + sparse.
Pay attention to:
- positivity violations and dist of \(A|X\)
- percent of outcome variance explainable by \(\mu\) (signal-to-noise ratio)
- percent of variance of \(\mu\) explainable by best linear approximation
Evidence for Goal 1
Across the board, AIPW and TMLE behave similarly in terms of point estimate and SE estimate in large samples when they have access to the same learners
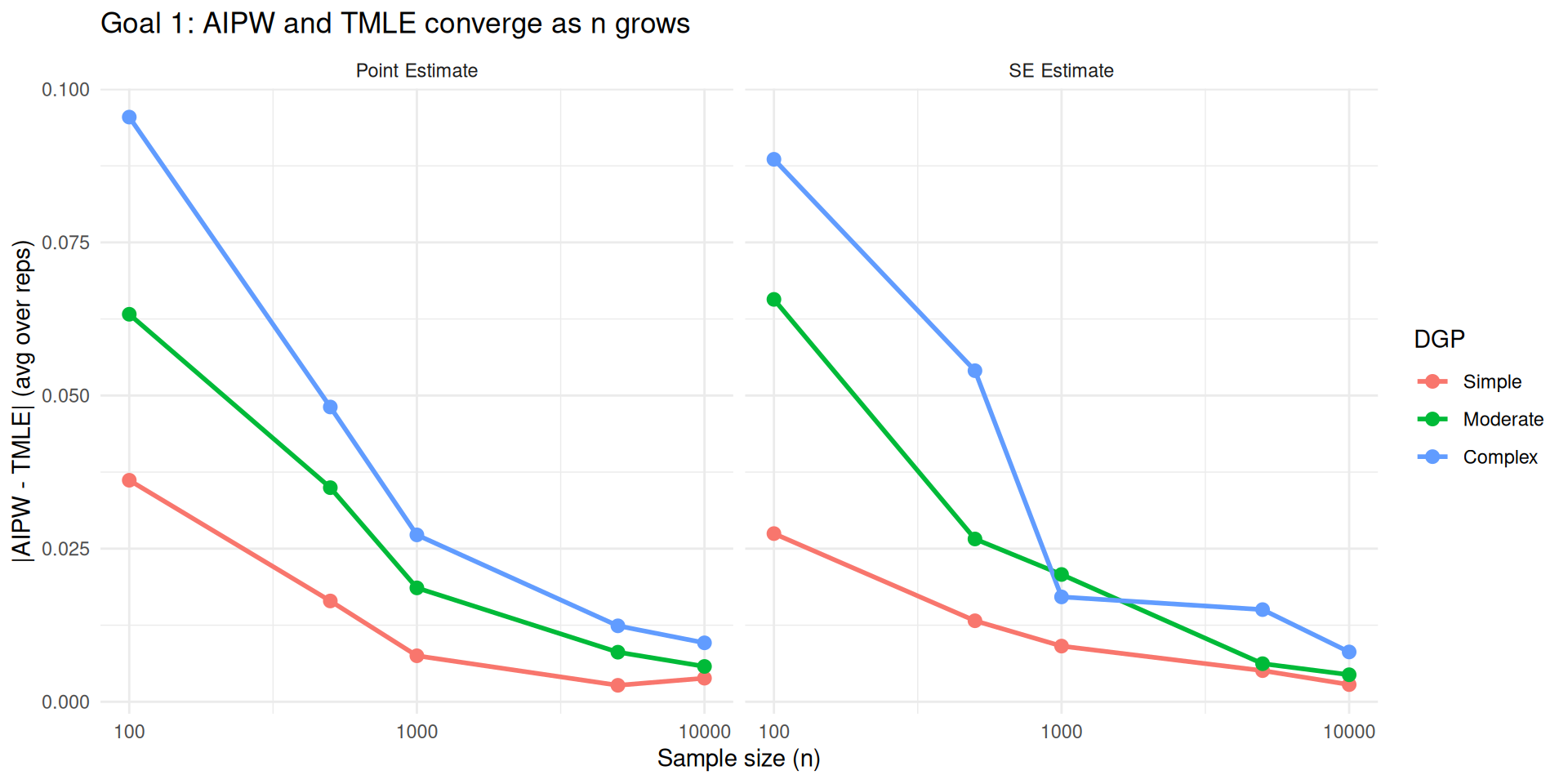
Settings for Goal 1
Across the board, AIPW and TMLE behave similarly in terms of point estimate and SE estimate in large samples when they have access to the same learners
- What DGPs? What estimand(s)? What learners?
- Do ~100 reps first, just get the vibe (do 2 reps first to see if code even runs!)
- \(n\) from 100 to 10k? Start with 3–4 increments, add more later
- Maybe 3 DGPs of increasing complexity/confounding. Simple \(W,A,Y\) setup w/ binary \(A\) for intelligibility. Doesn’t matter much for this goal
- For convenience, choose estimands that are computationally easy and work with the same DGP, e.g. ATE and incremental pscore. Having 2 estimands helps enforce the top-level generality claim, but stick to ATE for first round of output
- Learners: fast nonparametric learners for both OR and PS
Advice for Simulation Code
- Separate DGPs are separate objects with
drawmethod - Estimators are functions, separate, but can share innards
- separate out learning of common nuisances
- Generally return
(est, est_se) - Clear claims will help debug!
- Simulation loop over DGPs, estimators, and reps is a simple nested loop that produces a tidy data frame of results
- Use
furrrfor drop-in parallelization frommapin R, parallelize across reps
- Use
- Keep DGPs (and helpers) in one file, estimators (and helpers) in another, and run your sims/tests in a notebook, saving results to disk as needed
DGPs: Simplest Version
DGPs: More Modular
DGPs: OOP Approach
simple_dgp <- structure(
list(
rho = \(w1, w2) w1 + w2,
mu = \(a, w1, w2) w1 + w2 + a,
sigma = 1
),
class = "dgp"
)
complex_dgp <- structure(
list(
rho = \(w1, w2) w1 + w2 + w1 * abs(w2),
mu = \(a, w1, w2) w1 + w2 + abs(w2) + 0.5 * w1 * w2,
sigma = 1
),
class = "dgp"
)
draw <- function(x, ...) UseMethod("draw")
draw.dgp <- \(dgp, n) dgp %$% tibble(
W1 = runif(n, -1, 1),
W2 = runif(n, -1, 1),
A = rbinom(n, 1, plogis(rho(W1, W2))),
Y = mu(A, W1, W2) + rnorm(n, 0, sigma)
)
draw_intervene <- function(x, ...) UseMethod("draw_intervene")
draw_intervene.dgp <- \(dgp, n, treatment) dgp %$% tibble(
W1 = runif(n, -1, 1),
W2 = runif(n, -1, 1),
A = treatment(W1, W2),
Y = mu(A, W1, W2) + rnorm(n)
)DGPs: Usage
# A tibble: 3 × 4
W1 W2 A Y
<dbl> <dbl> <int> <dbl>
1 0.351 0.133 1 0.780
2 0.966 0.699 1 1.59
3 0.519 -0.621 1 0.846# A tibble: 3 × 4
W1 W2 A Y
<dbl> <dbl> <int> <dbl>
1 0.439 0.0288 1 1.26
2 -0.984 -0.997 0 -1.22
3 -0.249 0.163 1 -1.31# A tibble: 3 × 4
W1 W2 A Y
<dbl> <dbl> <dbl> <dbl>
1 0.335 0.866 1 1.77
2 -1.000 0.851 1 1.51
3 -0.583 0.468 1 1.21DGP Helpers
ATE <- function(x, ...) UseMethod("ATE")
ATE.dgp <- \(dgp, n = 1e6) {
mu0 <- dgp |> draw_intervene(n, \(w1, w2) 0) %$% mean(Y)
mu1 <- dgp |> draw_intervene(n, \(w1, w2) 1) %$% mean(Y)
mu1 - mu0
}
R2 <- function(x, ...) UseMethod("R2")
R2.dgp <- \(dgp, n = 1e6) {
pop <- dgp |>
draw(n) |>
mutate(mu = dgp$mu(A, W1, W2))
mu_lin <- lm(mu ~ A + W1 + W2, data = pop)
pop <- pop |>
mutate(mu_lin = predict(mu_lin, pop))
list(
R2 = pop %$% { var(mu) / var(Y) },
R2_lin = pop %$% { var(mu_lin) / var(mu) }
)
}Estimators: General Structure
So that it can be used like this:
- write code for simulation, not for a human user
Estimator: Difference in Means
Estimators: TMLE & AIPW
library(zeallot)
tmle <- function(data, nuisances) {
c(pi, alpha, mu) %<-% nuisances
eps <- lm(Y - mu(A, W1, W2) ~ alpha(A, W1, W2) - 1, data) |> coef() |> unname()
data |>
mutate(
mu1 = mu(1, W1, W2) + eps * alpha(1, W1, W2),
mu0 = mu(0, W1, W2) + eps * alpha(0, W1, W2),
phi = alpha(A, W1, W2) * (Y - mu(A, W1, W2)) + (mu1 - mu0)
) %$%
list(
est = mean(mu1 - mu0),
est_se = sd(phi) / sqrt(nrow(data))
)
}Learning Nuisances
library(tidymodels)
fit_learners <- function(data) {
# Convert A to factor for classification
data <- data |> mutate(A = factor(A))
pi_obj <- mars(prod_degree = 3, num_terms = 50) |>
set_engine("earth") |>
set_mode("classification") |>
fit(A ~ W1 + W2, data)
mu_obj <- mars(prod_degree = 3, num_terms = 50) |>
set_engine("earth") |>
set_mode("regression") |>
fit(Y ~ A + W1 + W2, data)
pi = \(w1, w2) predict(pi_obj, tibble(W1 = w1, W2 = w2), type = "prob")$.pred_1
alpha = \(a, w1, w2) a / pi(w1, w2) -
(1 - a) / (1 - pi(w1, w2))
mu = \(a, w1, w2) predict(mu_obj, tibble(A = factor(a, levels = c(0, 1)), W1 = w1, W2 = w2))$.pred
list(pi = pi, alpha = alpha, mu = mu)
}Sample Splitting
Simulation Loop
REPS <- 20
N <- 500
DGPS <- list(simple = simple_dgp, complex = complex_dgp)
ESTIMATORS <- list(dm = diff_means, tmle = tmle, aipw = aipw)
1:REPS |> map(\(rep) {
DGPS |> imap(\(dgp, dgp_name) {
nuisances <- dgp |> draw(N) |> fit_learners()
data <- dgp |> draw(N)
ESTIMATORS |> imap(\(estimator, estimator_name) {
res <- estimator(data, nuisances)
tibble(
rep = rep,
DGP = dgp_name,
estimator = estimator_name,
est = res$est,
se = res$est_se
)
}) |> bind_rows()
}) |> bind_rows()
}) |> bind_rows() -> results- Save the raw
resultsobject to file, possibly with a pointer to the commit hash for replicability - If you optimize runtime, you don’t have to worry so much about saving carefully since it’s so fast to regenerate!
Parallelize with furrr
library(furrr)
plan(multicore, workers = 10)
1:REPS |> furrr::future_map(\(rep) {
DGPS |> imap(\(dgp, dgp_name) {
nuisances <- dgp |> draw(N) |> fit_learners()
data <- dgp |> draw(N)
ESTIMATORS |> imap(\(estimator, estimator_name) {
res <- estimator(data, nuisances)
tibble(
rep = rep,
DGP = dgp_name,
estimator = estimator_name,
est = res$est,
se = res$est_se
)
}) |> bind_rows()
}) |> bind_rows()
}) |> bind_rows() -> results- before doing this, test with
REPS = 5and possibly fewer DGPs and estimators. - use
future_mapinstead ofmapand set up a parallel plan withplan()
Analysis
true_ATEs <- DGPS |> map_dbl(ATE)
results |>
group_by(DGP, estimator) |>
summarize(
bias = mean(est - true_ATEs[DGP]),
true_se = sd(est),
mean_est_se = mean(se),
rmse = sqrt(mean((est - true_ATEs[DGP])^2)),
coverage = mean(
(est - 1.96 * se <= true_ATEs[DGP]) &
(true_ATEs[DGP] <= est + 1.96 * se)
)
)# A tibble: 6 × 7
# Groups: DGP [2]
DGP estimator bias true_se mean_est_se rmse coverage
<chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 complex aipw -0.0137 0.0884 0.116 0.0873 1
2 complex dm 0.663 0.105 0.115 0.671 0
3 complex tmle -0.00608 0.0742 0.116 0.0726 1
4 simple aipw -0.0874 0.430 0.175 0.428 0.9
5 simple dm 0.594 0.130 0.112 0.608 0
6 simple tmle -0.00738 0.103 0.175 0.101 0.9From here you can use tidyr, kable, and ggplot to make tables and figures.
Tooling Demo
- Positron (files + notebook)
- GitHub Copilot in Positron Assistant
What About Documentation & Testing?
- Don’t bother, you’ll have to rewrite it all tomorrow
- Simple comments here and there suffice
- When you write a new function, unit test it in a Jupyter notebook a few times informally
- GitHub? Sure. Just commit to main, don’t worry about branches, PRs, etc.
Iteration
Problem!
In your initial TMLE simulation, you find that the ML plug-in estimator outperforms TMLE and AIPW in terms of MSE. Debiasing improves bias, but at cost of variance.
- Why is this bad for your claim?
- What do you do?
Solution
- having a clear goal made it easy to know if result was “good” or “bad”
- understand theory: why is debiasing necessary?
- modify claim:
Like AIPW, TMLE improves the MSE of plugins in moderate sample sizes when overlap is reasonable and initial estimates are biased and allows for good coverage of Wald intervals based on IF SEs.
- good claims are not overstated!
- edge cases clarify claims: design sims to archetype extremes
Conclusions
How to do Simulations
Top Tips
🎯 Thoughtful Goals = Direction!
- practical utility
- key insight
- audience
📊 Communication = Focus!
- structure + organize
- minimize cognitive load
⚙️ Clean, Fast Sims = Productivity!
- simplest DGP
- minimize compute
- modular code